Bringing Light to Extracellular Vesicleshristopher Pratt, hD
| Ver aquí ExoBrite™ EV Membrane Staining Kits |
Origin of extracellular vesicles
Extracellular vesicles (EVs) are heterogeneous, membrane-encapsulated particles that are secreted by cells across the tree of life.1–3 Though originally thought of as strictly a waste disposal system, EVs have been shown to have myriad cell communication functions. Two main classes of EVs exist: microvesicles are large (200-2000 nm diameter) vesicles produced by direct budding from the plasma membrane (PM), while exosomes are small (40-200 nm) and are primarily produced when a multivesicular body fuses with the PM, releasing intraluminal vesicles as exosomes. Exosomes directly budding from the PM has been observed in T-cells; however, the frequency of this phenomenon in other cell types is unclear.2,4
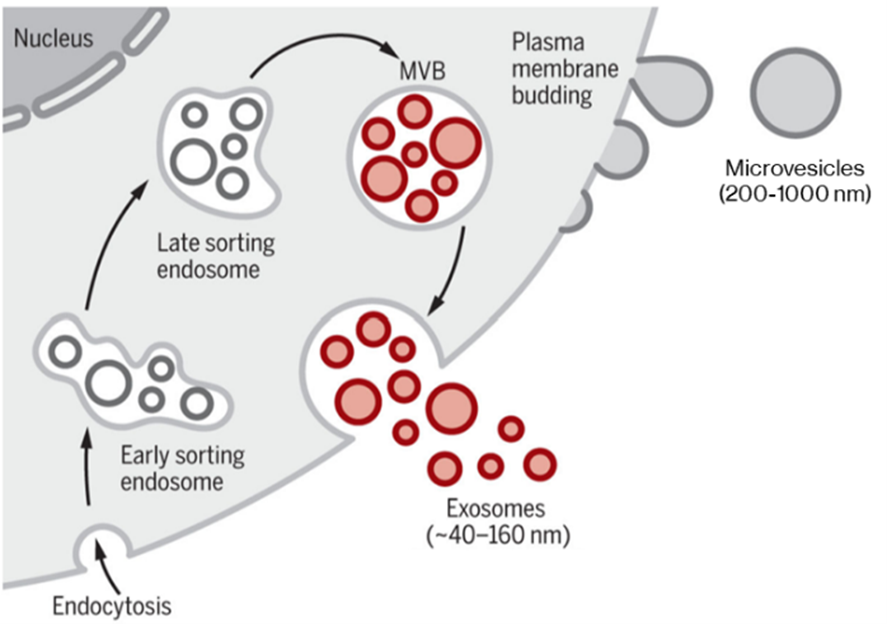
Classical modes of EV generation, producing microvesicles directly from plasma membrane (PM) and exosomes via the late sorting endosome and multivesicular body (MVB). Certain cell types can also produce exosomes by direct PM budding. Adapted from Kalluri & LeBleu (2020).5 Reproduced under the Creative Commons license.
Exosomes and microvesicles share several features; their proteomes resemble the originating cell but are enriched in certain proteins. Tetraspanins are membrane proteins that act as molecular organizers and include CD9 and CD81 in microvesicles and exosomes, while CD63 is a specific marker for exosomes.1,2 Exosomes also contain a dense intraluminal core of scaffolding and aggregated protein bound with chaperones, primarily HSP70.2 RNA and DNA are common cargoes in EVs and can elicit responses including gene transcription or suppression in recipient cells.1,2,5,6
The tropism of EVs varies by their composition and the recipient cell type: the tetraspanins and other membrane proteins determine EV specificity by interacting with integrins, lectins, or extracellular matrix components.1 Exosomes secreted by cortical neurons are taken up only by neurons, whereas exosomes from the N2a neuroblastoma cell line target neurons and glia.7 The mechanism of EV uptake –macropinocytosis, phagocytosis, clathrin or caveolin-dependent endocytosis, or direct membrane fusion—is also cell-type dependent.1
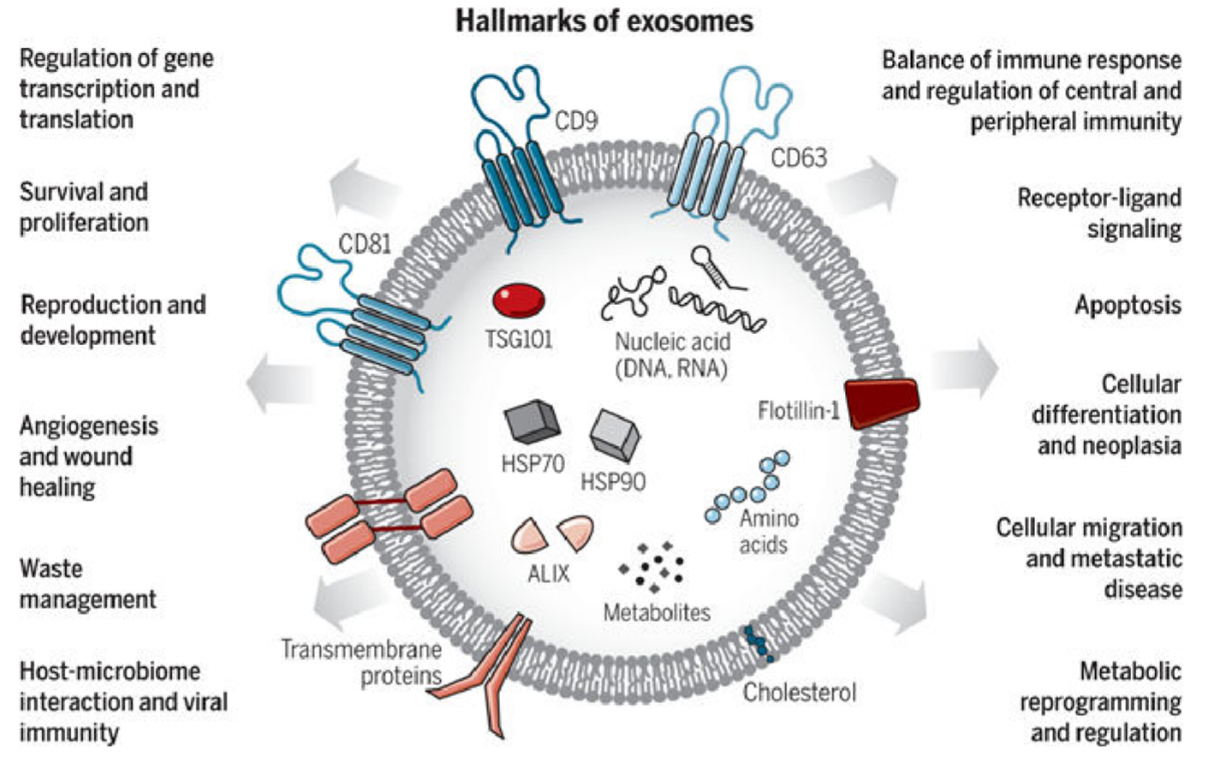
Exosomes are varied in size and composition, carrying diverse cargoes including membrane proteins (particularly CD63 and CD81). Exosomes are involved in a variety of functions. From Kalluri & LeBleu (2020).5 Reproduced under the Creative Commons license.
Extracellular vesicles in disease and diagnostics
EVs carry cargoes of proteins, RNA, and DNA that function in the recipient cells.1,2,5,6 Microvesicles secreted by cancer cells (sometimes called oncosomes) can transport oncogenic mRNA that promotes invasion and metastasis in breast cancer cells,8 and exosomes from pancreatic cancer cells initiate transformation in normal cells.9 EVs can also contribute to immunity and immunosuppression: antigen presenting cells secrete exosomes containing major histocompatibility complex class II bound with antigen, which can promote adaptive immunity over a distance,5 and tumor-derived EVs can promote immunosuppression within the tumor microenvironment.5 Viruses can use EVs to escape from cells and infect others; for example, rotavirus capsids hide within large secreted microvesicles and HIV Gag proteins use T-cell derived exosomes.4,10
Exosomes have also been associated with neurodegeneration. Prion protein is present in brain-derived exosomes, where it plays roles in immunomodulation, plasticity, and normal brain function. Pathogenic prion protein in spongiform encephalopathies also gets carried by exosomes along with other factors—microRNAs in particular—that potentiate its neurotoxic effects.11 MicroRNAs associated with the progression of Alzheimer’s Disease and Parkinson’s Disease have also been found in brain-derived exosomes. Exosomes’ roles in the pathogenesis of these and other neurodegenerative diseases remain to be determined.
Liquid Biopsies
Because EVs can travel far and wide through biofluids, one of the most exciting prospects is developing non-invasive “liquid biopsies” to monitor for disease or to sample hard-to-access organs such as the brain or pancreas.6,12,13 Glypican-1 is seen in pancreatic cancer exosomes isolated from the blood of patients and can be used as a biomarker.14 Liquid biopsies could be used for early detection of cancers, neurodegenerative disease, pregnancy disorders, and other applications.12
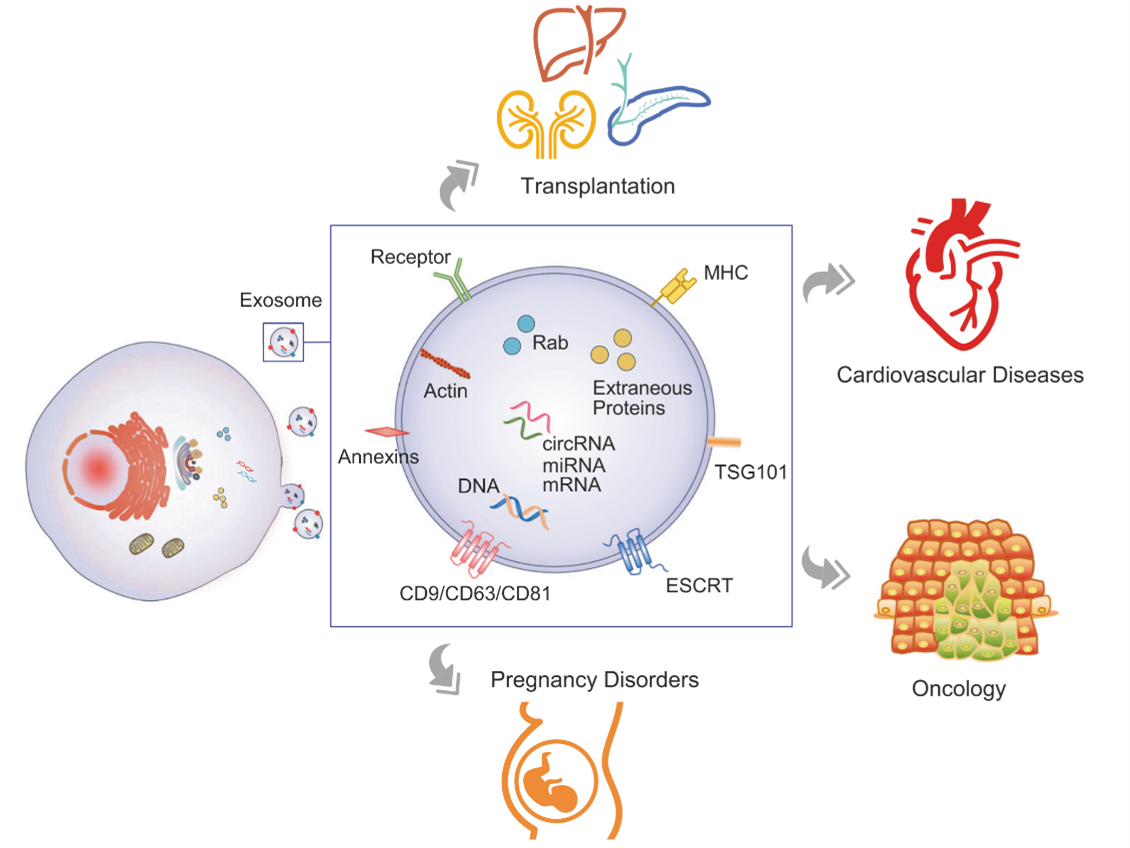
Circulating EVs make non-invasive liquid biopsies possible. Figure from Zhou et al. 2020.12 Reproduced under the Creative Commons license.
The challenges of analyzing EVs
As small heterogeneous extracellular particles, EVs pose challenges to characterize; a principal confounder is the lack of standardized isolation and analysis methods.5,15,16 EVs have classically been isolated by ultracentrifugation or sucrose gradient ultracentrifugation,2,3 but the purity of isolated EVs is poor.15 New isolation techniques such as acoustic nanofiltering and microfluidic immuno-isolation improve purity, but require specialized equipment.
Scanning and transmission electron microscopy can readily visualize EVs, but sample preparation creates artifacts and, depending on the preparation method, can cause EV sampling bias.2,4,16 Fluorescence imaging with lipophilic dyes in culture systems are confounded by other small particles in biofluids and cell culture media like serum proteins and lipoproteins.17 Mass spectroscopy, NGS and proteomics are commonly used to analyze EV content, but heterogeneity and purity are lingering concerns.
Analyzing exosomes with flow cytometry is a new field; one challenge is distinguishing exosomes from cellular debris. Fortunately, EVs have well-defined markers and properties that can be exploited:1,16 CD9, CD63, and CD81 are markers for EVs, with CD63 being specific to exosomes. In certain types of EVs, Ep-CAM and HSP70 are useful targets.1,5 For flow cytometry of EVs, it is crucial to avoid forming aggregates as the loss of individual resolution is disastrous for such a heterogeneous population.15 Antibodies validated for EV detection by flow cytometry are a key starting point. Membrane stains validated for flow cytometry are useful for measuring EV size by fluorescence intensity.
The Future of EVs
EVs represent an exciting and booming field of cell biology with life-changing diagnostic and therapeutic potential. Better understanding and characterization of EVs is critical in making this potential into reality. Biotium’s curated EV technology resource will assemble the latest validated tools for EV research. Without a doubt, EV research will drive advances in cell biology and medicine.
References
- van Niel, G., D’Angelo, G. & Raposo, G. Shedding light on the cell biology of extracellular vesicles. Nat Rev Mol Cell Biol 19, 213–228 (2018). DOI: 10.1038/nrm.2017.125
- Pegtel, D. M. & Gould, S. J. Exosomes. Annu Rev Biochem 88, 487–514 (2019). DOI: 10.1146/annurev-biochem-013118-111902
- Trams, E. G., Lauter, C. J., Salem, N. & Heine, U. Exfoliation of membrane ecto-enzymes in the form of micro-vesicles. Biochim Biophys Acta 645, 63–70 (1981). DOI: 10.1016/0005-2736(81)90512-5
- Booth, A. M. et al. Exosomes and HIV Gag bud from endosome-like domains of the T cell plasma membrane. J Cell Biol 172, 923–935 (2006). DOI: 10.1083/jcb.200508014
- Kalluri, R. & LeBleu, V. S. The biology, function, and biomedical applications of exosomes. Science 367, (2020). DOI: 10.1126/science.aau6977
- Armstrong, D. & Wildman, D. E. Extracellular Vesicles and the Promise of Continuous Liquid Biopsies. J Pathol Transl Med 52, 1–8 (2018). DOI: 10.4132/jptm.2017.05.21
- Laulagnier, K. et al. Amyloid precursor protein products concentrate in a subset of exosomes specifically endocytosed by neurons. Cell. Mol. Life Sci. 75, 757–773 (2018). DOI: 10.1007/s00018-017-2664-0
- Le, M. T. N. et al. miR-200-containing extracellular vesicles promote breast cancer cell metastasis. J Clin Invest 124. DOI: 10.1172/JCI75695
- Stefanius, K. et al. Human pancreatic cancer cell exosomes, but not human normal cell exosomes, act as an initiator in cell transformation. Elife 8, (2019). DOI: 10.7554/eLife.40226
- Santiana, M. et al. Vesicle-Cloaked Virus Clusters Are Optimal Units for Inter-organismal Viral Transmission. Cell Host & Microbe 24, 208-220.e8 (2018). DOI: 10.1016/j.chom.2018.07.006
- Song, Z. et al. Brain Derived Exosomes Are a Double-Edged Sword in Alzheimer’s Disease. Front. Mol. Neurosci. 13, (2020). DOI: 10.3389/fnmol.2020.00079
- Zhou, B. et al. Application of exosomes as liquid biopsy in clinical diagnosis. Signal Transduction and Targeted Therapy 5, 1–14 (2020). DOI: 10.1038/s41392-020-00258-9
- Coleman, B. M. & Hill, A. F. Extracellular vesicles – Their role in the packaging and spread of misfolded proteins associated with neurodegenerative diseases. Seminars in Cell & Developmental Biology 40, 89–96 (2015). DOI: 10.1016/j.semcdb.2015.02.007
- Melo, S. A. et al. Glypican-1 identifies cancer exosomes and detects early pancreatic cancer. Nature 523, 177–182 (2015). DOI: 10.1038/nature14581
- Doyle, L. M. & Wang, M. Z. Overview of Extracellular Vesicles, Their Origin, Composition, Purpose, and Methods for Exosome Isolation and Analysis. Cells 8, (2019),. DOI: 10.3390/cells8070727
- Shao, H. et al. New Technologies for Analysis of Extracellular Vesicles. Chem Rev 118, 1917–1950 (2018).DOI: 10.1021/acs.chemrev.7b00534
- Takov, K., Yellon, D. M. & Davidson, S. M. Confounding factors in vesicle uptake studies using fluorescent lipophilic membrane dyes. J Extracell Vesicles 6, (2017). DOI: 10.1080/20013078.2017.1388731
Christopher Pratt, PhD 

Christopher Pratt earned his PhD in cell biology and neuroscience from Carnegie Mellon University in 2016 under the guidance of Marcel Bruchez, PhD. For his thesis, he developed and applied novel fluorescent probes to understand synaptic vesicle and ion channel trafficking in the brain, making him a ardent microscopist. With a passion for communicating science and data, Christopher is a regular contributor to Biotium’s Full Spectrum Blog and is a freelance science and medical writer, analyst, and research consultant based in Chicago, Illinois. Visit his website, twitter, or LinkedIn and get in touch!
Biotium Source

